2.2. 高级NumPy ¶
Author: Pauli Virtanen
NumPy is at the base of Python’s scientific stack of tools. 它的目的是对一块内存中的许多元素实现高效的操作。理解它如何工作的细节有助于有效地利用其灵活性、采用有用的快捷操作。
This section covers:
- NumPy数组的剖析及其结果。提示和技巧。
- 通用函数:什么是通用函数、为什么使用通用函数、以及如果你想要一个新的通用函数要做什么。
- 与其他工具集成:NumPy提供了几种方法来在ndarray中包装任何数据,没有不必要的拷贝。
- Recently added features, and what’s in them: PEP 3118 buffers, generalized ufuncs, ...
Prerequisites
- NumPy
- Cython
- Pillow(Python图像库,在几个例子中用到)
Chapter contents
Tip
In this section, numpy will be imported as follows:
>>> import numpy as np
2.2.1. Life of ndarray¶
2.2.1.1. It’s...¶
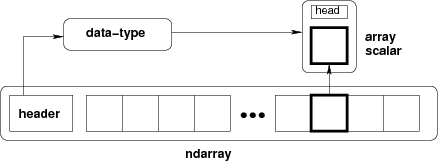
ndarray =
内存块 + 索引方案 + 数据类型描述符
- raw data
- how to locate an element
- how to interpret an element
typedef struct PyArrayObject {
PyObject_HEAD
/* Block of memory */
char *data;
/* Data type descriptor */
PyArray_Descr *descr;
/* Indexing scheme */
int nd;
npy_intp *dimensions;
npy_intp *strides;
/* Other stuff */
PyObject *base;
int flags;
PyObject *weakreflist;
} PyArrayObject;
2.2.1.2. Block of memory¶
>>> x = np.array([1, 2, 3], dtype=np.int32)
>>> x.data
<... at ...>
>>> str(x.data)
'\x01\x00\x00\x00\x02\x00\x00\x00\x03\x00\x00\x00'
Memory address of the data:
>>> x.__array_interface__['data'][0]
64803824
The whole __array_interface__:
>>> x.__array_interface__
{'data': (35828928, False),
'descr': [('', '<i4')],
'shape': (4,),
'strides': None,
'typestr': '<i4',
'version': 3}
Reminder: two ndarrays may share the same memory:
>>> x = np.array([1, 2, 3, 4])
>>> y = x[:-1]
>>> x[0] = 9
>>> y
array([9, 2, 3])
Memory does not need to be owned by an ndarray:
>>> x = b'1234' # The 'b' is for "bytes", necessary in Python 3
x是一个字符串(在Python 3中是一个字节),我们可以将其数据表示为int数组:
>>> y = np.frombuffer(x, dtype=np.int8)
>>> y.data
<... at ...>
>>> y.base is x
True
>>> y.flags
C_CONTIGUOUS : True
F_CONTIGUOUS : True
OWNDATA : False
WRITEABLE : False
ALIGNED : True
UPDATEIFCOPY : False
owndata和writeable标志指示内存块的状态。
2.2.1.3. Data types¶
2.2.1.3.1. 描述符¶
dtype describes a single item in the array:
| type | 数据的标量类型,下面中的一个: int8、int16、float64等。 (fixed size) str、unicode、void(大小可变) |
| itemsize | size of the data block |
| byteorder | 字节顺序: 大端序 > / 小端序 < / 不可用| |
| fields | sub-dtypes, if it’s a structured data type |
| shape | 如果是一个子数组,则它为数组的形状 |
>>> np.dtype(int).type
<type 'numpy.int64'>
>>> np.dtype(int).itemsize
8
>>> np.dtype(int).byteorder
'='
2.2.1.3.2. 示例:读取.wav文件¶
The .wav file header:
| chunk_id | "RIFF" |
| chunk_size | 4-byte unsigned little-endian integer |
| format | "WAVE" |
| fmt_id | "fmt " |
| fmt_size | 4-byte unsigned little-endian integer |
| audio_fmt | 2-byte unsigned little-endian integer |
| num_channels | 2-byte unsigned little-endian integer |
| sample_rate | 4-byte unsigned little-endian integer |
| byte_rate | 4-byte unsigned little-endian integer |
| block_align | 2-byte unsigned little-endian integer |
| bits_per_sample | 2-byte unsigned little-endian integer |
| data_id | "data" |
| data_size | 4-byte unsigned little-endian integer |
- 44-byte block of raw data (in the beginning of the file)
- ... followed by
data_sizebytes of actual sound data.
.wav文件头作为NumPy结构化数据类型:
>>> wav_header_dtype = np.dtype([
... ("chunk_id", (bytes, 4)), # flexible-sized scalar type, item size 4
... ("chunk_size", "<u4"), # little-endian unsigned 32-bit integer
... ("format", "S4"), # 4-byte string
... ("fmt_id", "S4"),
... ("fmt_size", "<u4"),
... ("audio_fmt", "<u2"), #
... ("num_channels", "<u2"), # .. more of the same ...
... ("sample_rate", "<u4"), #
... ("byte_rate", "<u4"),
... ("block_align", "<u2"),
... ("bits_per_sample", "<u2"),
... ("data_id", ("S1", (2, 2))), # sub-array, just for fun!
... ("data_size", "u4"),
... #
... # the sound data itself cannot be represented here:
... # it does not have a fixed size
... ])
另见
wavreader.py
>>> wav_header_dtype['format']
dtype('S4')
>>> wav_header_dtype.fields
dict_proxy({'block_align': (dtype('uint16'), 32), 'format': (dtype('S4'), 8), 'data_id': (dtype(('S1', (2, 2))), 36), 'fmt_id': (dtype('S4'), 12), 'byte_rate': (dtype('uint32'), 28), 'chunk_id': (dtype('S4'), 0), 'num_channels': (dtype('uint16'), 22), 'sample_rate': (dtype('uint32'), 24), 'bits_per_sample': (dtype('uint16'), 34), 'chunk_size': (dtype('uint32'), 4), 'fmt_size': (dtype('uint32'), 16), 'data_size': (dtype('uint32'), 40), 'audio_fmt': (dtype('uint16'), 20)})
>>> wav_header_dtype.fields['format']
(dtype('S4'), 8)
- 第一个元素是结构化数据中的子类型,对应于名称
format - 第二个是它从元素开始的偏移量(以字节为单位)
练习
迷你练习,通过使用偏移量生成一个“稀疏”的dtype,并只有一些字段:
>>> wav_header_dtype = np.dtype(dict(
... names=['format', 'sample_rate', 'data_id'],
... offsets=[offset_1, offset_2, offset_3], # counted from start of structure in bytes
... formats=list of dtypes for each of the fields,
... ))
and use that to read the sample rate, and data_id (as sub-array).
>>> f = open('data/test.wav', 'r')
>>> wav_header = np.fromfile(f, dtype=wav_header_dtype, count=1)
>>> f.close()
>>> print(wav_header)
[ ('RIFF', 17402L, 'WAVE', 'fmt ', 16L, 1, 1, 16000L, 32000L, 2, 16, [['d', 'a'], ['t', 'a']], 17366L)]
>>> wav_header['sample_rate']
array([16000], dtype=uint32)
Let’s try accessing the sub-array:
>>> wav_header['data_id']
array([[['d', 'a'],
['t', 'a']]],
dtype='|S1')
>>> wav_header.shape
(1,)
>>> wav_header['data_id'].shape
(1, 2, 2)
当访问子数组时,维度被添加到末尾!
Note
有已经存在的模块,如wavfile、audiolab等用于加载声音数据...
2.2.1.3.3. 转换和重新解释/视图¶
转换
- 在赋值的时候
- 在数组构造的时候
- 在算术计算的时候
- etc.
- 以及手动转换:
.astype(dtype)
data re-interpretation
- 手动去做:
.view(dtype)
2.2.1.3.3.1. Casting¶
算术计算中的转换,简而言之:
- only type (not value!) of operands matters
- 选择能够代表两者的最大“安全”类型
- scalars can “lose” to arrays in some situations
通常拷贝数据时的转换:
>>> x = np.array([1, 2, 3, 4], dtype=np.float) >>> x array([ 1., 2., 3., 4.]) >>> y = x.astype(np.int8) >>> y array([1, 2, 3, 4], dtype=int8) >>> y + 1 array([2, 3, 4, 5], dtype=int8) >>> y + 256 array([257, 258, 259, 260], dtype=int16) >>> y + 256.0 array([ 257., 258., 259., 260.]) >>> y + np.array([256], dtype=np.int32) array([257, 258, 259, 260], dtype=int32)
在setitem时的类型转换:在元素赋值时数组的dtype不更改:
>>> y[:] = y + 1.5 >>> y array([2, 3, 4, 5], dtype=int8)
Note
Exact rules: see numpy documentation
2.2.1.3.3.2. 重新解释/视图¶
Data block in memory (4 bytes)
0x01|| 0x02|| 0x03|| 0x04- 4个uint8,或者
- 4个int8,或者
- 2个int16,或者
- 1个int32,或者
- 1个float32,或者
- ...
How to switch from one to another?
Switch the dtype:
>>> x = np.array([1, 2, 3, 4], dtype=np.uint8) >>> x.dtype = "<i2" >>> x array([ 513, 1027], dtype=int16) >>> 0x0201, 0x0403 (513, 1027)
0x010x02|| 0x030x04Note
little-endian:在内存中,最低有效字节位于左边
Create a new view:
>>> y = x.view("<i4") >>> y array([67305985], dtype=int32) >>> 0x04030201 67305985
0x010x020x030x04
Note
.view()生成视图,不复制(或更改)内存块only changes the dtype (and adjusts array shape):
>>> x[1] = 5 >>> y array([328193], dtype=int32) >>> y.base is x True
Mini-exercise: data re-interpretation
另见
view-colors.py
You have RGBA data in an array:
>>> x = np.zeros((10, 10, 4), dtype=np.int8)
>>> x[:, :, 0] = 1
>>> x[:, :, 1] = 2
>>> x[:, :, 2] = 3
>>> x[:, :, 3] = 4
where the last three dimensions are the R, B, and G, and alpha channels.
如何做一个字段名称为'r','g','b','a'的(10, 10)结构化数组而不复制数据?
>>> y = ...
>>> assert (y['r'] == 1).all()
>>> assert (y['g'] == 2).all()
>>> assert (y['b'] == 3).all()
>>> assert (y['a'] == 4).all()
答案
Warning
另一个正好占用4个字节内存的数组:
>>> y = np.array([[1, 3], [2, 4]], dtype=np.uint8).transpose()
>>> x = y.copy()
>>> x
array([[1, 2],
[3, 4]], dtype=uint8)
>>> y
array([[1, 2],
[3, 4]], dtype=uint8)
>>> x.view(np.int16)
array([[ 513],
[1027]], dtype=int16)
>>> 0x0201, 0x0403
(513, 1027)
>>> y.view(np.int16)
array([[ 769, 1026]], dtype=int16)
- What happened?
- ... we need to look into what
x[0,1]actually means
>>> 0x0301, 0x0402
(769, 1026)
2.2.1.4. Indexing scheme: strides¶
2.2.1.4.1. 要点¶
The question:
>>> x = np.array([[1, 2, 3],
... [4, 5, 6],
... [7, 8, 9]], dtype=np.int8)
>>> str(x.data)
'\x01\x02\x03\x04\x05\x06\x07\x08\t'
At which byte in ``x.data`` does the item ``x[1, 2]`` begin?
The answer (in NumPy)
- strides:找到下一个元素跳转的字节数
- 每个维度一个stride
>>> x.strides
(3, 1)
>>> byte_offset = 3*1 + 1*2 # to find x[1, 2]
>>> x.flat[byte_offset]
6
>>> x[1, 2]
6
- simple, **flexible**
2.2.1.4.1.1. C和Fortran的顺序¶
>>> x = np.array([[1, 2, 3],
... [4, 5, 6]], dtype=np.int16, order='C')
>>> x.strides
(6, 2)
>>> str(x.data)
'\x01\x00\x02\x00\x03\x00\x04\x00\x05\x00\x06\x00'
- Need to jump 6 bytes to find the next row
- Need to jump 2 bytes to find the next column
>>> y = np.array(x, order='F')
>>> y.strides
(2, 4)
>>> str(y.data)
'\x01\x00\x04\x00\x02\x00\x05\x00\x03\x00\x06\x00'
- Need to jump 2 bytes to find the next row
- Need to jump 4 bytes to find the next column
更高的维度类似:
- C:最后的维度变化最快(=较小的stride)
- F:第一个维度变化最快
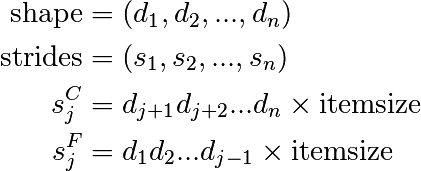
Note
Now we can understand the behavior of .view():
>>> y = np.array([[1, 3], [2, 4]], dtype=np.uint8).transpose()
>>> x = y.copy()
转置不影响数据的内存布局,只影响stride
>>> x.strides
(2, 1)
>>> y.strides
(1, 2)
>>> str(x.data)
'\x01\x02\x03\x04'
>>> str(y.data)
'\x01\x03\x02\x04'
- 当解释为2个int16时,结果不同
.copy()creates new arrays in the C order (by default)
2.2.1.4.1.2. 用整数切片¶
- 所有东西都可以通过只更改
shape、strides来表示,并可能只通过调整data指针表示! - 永远不会复制数据
>>> x = np.array([1, 2, 3, 4, 5, 6], dtype=np.int32)
>>> y = x[::-1]
>>> y
array([6, 5, 4, 3, 2, 1], dtype=int32)
>>> y.strides
(-4,)
>>> y = x[2:]
>>> y.__array_interface__['data'][0] - x.__array_interface__['data'][0]
8
>>> x = np.zeros((10, 10, 10), dtype=np.float)
>>> x.strides
(800, 80, 8)
>>> x[::2,::3,::4].strides
(1600, 240, 32)
类似地,转置永远不会生成拷贝(它只是交换stride):
>>> x = np.zeros((10, 10, 10), dtype=np.float) >>> x.strides (800, 80, 8) >>> x.T.strides (8, 80, 800)
但是:不是所有的reshape操作都可以通过使用stride来表示:
>>> a = np.arange(6, dtype=np.int8).reshape(3, 2)
>>> b = a.T
>>> b.strides
(1, 2)
So far, so good. However:
>>> str(a.data)
'\x00\x01\x02\x03\x04\x05'
>>> b
array([[0, 2, 4],
[1, 3, 5]], dtype=int8)
>>> c = b.reshape(3*2)
>>> c
array([0, 2, 4, 1, 3, 5], dtype=int8)
这里,只用一个stride和a的内存块,没有办法表示数组c。Therefore, the reshape operation needs to make a copy here.
2.2.1.4.2. Example: fake dimensions with strides¶
Stride操纵
>>> from numpy.lib.stride_tricks import as_strided
>>> help(as_strided)
as_strided(x, shape=None, strides=None)
Make an ndarray from the given array with the given shape and strides
Warning
as_strided不会检查是否在内存块边界内...
>>> x = np.array([1, 2, 3, 4], dtype=np.int16)
>>> as_strided(x, strides=(2*2, ), shape=(2, ))
array([1, 3], dtype=int16)
>>> x[::2]
array([1, 3], dtype=int16)
另见
stride-fakedims.py
Exercise
array([1, 2, 3, 4], dtype=np.int8) -> array([[1, 2, 3, 4], [1, 2, 3, 4], [1, 2, 3, 4]], dtype=np.int8)只使用
as_strided:Hint: byte_offset = stride[0]*index[0] + stride[1]*index[1] + ...
Spoiler
2.2.1.4.3. Broadcasting¶
- 用它做一些有用的东西:
[1, 2, 3, 4]和[5, 6, 7]的外积
>>> x = np.array([1, 2, 3, 4], dtype=np.int16)
>>> x2 = as_strided(x, strides=(0, 1*2), shape=(3, 4))
>>> x2
array([[1, 2, 3, 4],
[1, 2, 3, 4],
[1, 2, 3, 4]], dtype=int16)
>>> y = np.array([5, 6, 7], dtype=np.int16)
>>> y2 = as_strided(y, strides=(1*2, 0), shape=(3, 4))
>>> y2
array([[5, 5, 5, 5],
[6, 6, 6, 6],
[7, 7, 7, 7]], dtype=int16)
>>> x2 * y2
array([[ 5, 10, 15, 20],
[ 6, 12, 18, 24],
[ 7, 14, 21, 28]], dtype=int16)
... seems somehow familiar ...
>>> x = np.array([1, 2, 3, 4], dtype=np.int16)
>>> y = np.array([5, 6, 7], dtype=np.int16)
>>> x[np.newaxis,:] * y[:,np.newaxis]
array([[ 5, 10, 15, 20],
[ 6, 12, 18, 24],
[ 7, 14, 21, 28]], dtype=int16)
- Internally, array broadcasting is indeed implemented using 0-strides.
2.2.1.4.4. More tricks: diagonals¶
另见
stride-diagonals.py
Challenge
Pick diagonal entries of the matrix: (assume C memory order):
>>> x = np.array([[1, 2, 3], ... [4, 5, 6], ... [7, 8, 9]], dtype=np.int32) >>> x_diag = as_strided(x, shape=(3,), strides=(???,))Pick the first super-diagonal entries
[2, 6].And the sub-diagonals?
- (Hint to the last two: slicing first moves the point where striding
- starts from.)
答案
另见
stride-diagonals.py
Challenge
Compute the tensor trace:
>>> x = np.arange(5*5*5*5).reshape(5, 5, 5, 5) >>> s = 0 >>> for i in range(5): ... for j in range(5): ... s += x[j, i, j, i]by striding, and using
sum()on the result.>>> y = as_strided(x, shape=(5, 5), strides=(TODO, TODO)) >>> s2 = ... >>> assert s == s2
答案
2.2.1.4.5. CPU缓存的影响¶
Memory layout can affect performance:
In [1]: x = np.zeros((20000,))
In [2]: y = np.zeros((20000*67,))[::67]
In [3]: x.shape, y.shape
((20000,), (20000,))
In [4]: %timeit x.sum()
100000 loops, best of 3: 0.180 ms per loop
In [5]: %timeit y.sum()
100000 loops, best of 3: 2.34 ms per loop
In [6]: x.strides, y.strides
((8,), (536,))
Smaller strides are faster?
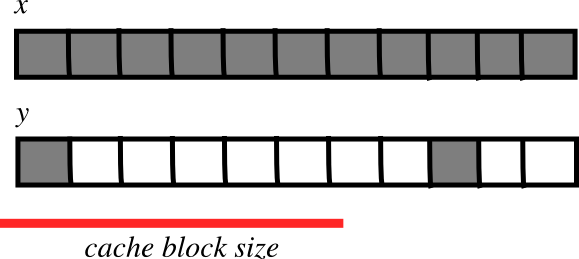
- CPU pulls data from main memory to its cache in blocks
- 如果连续操作的许多数组元素在一个单一的块内(stride较小):
 fewer transfers needed
fewer transfers needed faster
faster
另见
numexpr is designed to mitigate cache effects in array computing.
2.2.1.4.6. Example: inplace operations (caveat emptor)¶
Sometimes,
>>> a -= b
与下面的语句是不等同的
>>> a -= b.copy()
>>> x = np.array([[1, 2], [3, 4]])
>>> x -= x.transpose()
>>> x
array([[ 0, -1],
[ 4, 0]])
>>> y = np.array([[1, 2], [3, 4]])
>>> y -= y.T.copy()
>>> y
array([[ 0, -1],
[ 1, 0]])
xandx.transpose()share datax -= x.transpose()逐个元素修改数据...- 因为
x和x.transpose()有不同的striding,修改的数据重新出现在RHS
2.2.1.5. 剖析结果 ¶

- memory block: may be shared,
.base,.data - data type descriptor: structured data, sub-arrays, byte order, casting, viewing,
.astype(),.view() - strided indexing: strides, C/F-order, slicing w/ integers,
as_strided, broadcasting, stride tricks,diag, CPU cache coherence
2.2.2. 通用函数 ¶
2.2.2.1. What they are?¶
Ufunc对数组的所有元素执行元素级别操作。
Examples:
np.add, np.subtract, scipy.special.*, ...
自动支持:broadcasting、casting...
The author of an ufunc only has to supply the elementwise operation, NumPy takes care of the rest.
The elementwise operation needs to be implemented in C (or, e.g., Cython)
2.2.2.1.1. Parts of an Ufunc¶
Provided by user
void ufunc_loop(void **args, int *dimensions, int *steps, void *data) { /* * int8 output = elementwise_function(int8 input_1, int8 input_2) * * This function must compute the ufunc for many values at once, * in the way shown below. */ char *input_1 = (char*)args[0]; char *input_2 = (char*)args[1]; char *output = (char*)args[2]; int i; for (i = 0; i < dimensions[0]; ++i) { *output = elementwise_function(*input_1, *input_2); input_1 += steps[0]; input_2 += steps[1]; output += steps[2]; } }
The NumPy part, built by
char types[3] types[0] = NPY_BYTE /* type of first input arg */ types[1] = NPY_BYTE /* type of second input arg */ types[2] = NPY_BYTE /* type of third input arg */ PyObject *python_ufunc = PyUFunc_FromFuncAndData( ufunc_loop, NULL, types, 1, /* ntypes */ 2, /* num_inputs */ 1, /* num_outputs */ identity_element, name, docstring, unused)
- ufunc也可以支持多种不同输入—输出类型组合。
2.2.2.1.2. Making it easier¶
ufunc_loopis of very generic form, and NumPy provides pre-made onesPyUfunc_f_ffloat elementwise_func(float input_1)PyUfunc_ff_ffloat elementwise_func(float input_1, float input_2)PyUfunc_d_ddouble elementwise_func(double input_1)PyUfunc_dd_ddouble elementwise_func(double input_1, double input_2)PyUfunc_D_Delementwise_func(npy_cdouble *input, npy_cdouble* output)PyUfunc_DD_Delementwise_func(npy_cdouble *in1, npy_cdouble *in2, npy_cdouble* out)- Only
elementwise_funcneeds to be supplied - ... except when your elementwise function is not in one of the above forms
- Only
2.2.2.2. Exercise: building an ufunc from scratch¶
The Mandelbrot fractal is defined by the iteration
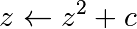
其中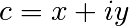 is a complex number. This iteration is repeated – if
is a complex number. This iteration is repeated – if 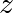 stays finite no matter how long the iteration runs,
stays finite no matter how long the iteration runs, 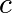 belongs to the Mandelbrot set.
belongs to the Mandelbrot set.
Make ufunc called
mandel(z0, c)that computes:z = z0 for k in range(iterations): z = z*z + c
例如迭代100次,或者直到
z.real**2 + z.imag**2 > 1000。Use it to determine which c are in the Mandelbrot set.Our function is a simple one, so make use of the
PyUFunc_*helpers.用Cython编写
另见
mandel.pyx, mandelplot.py
Reminder: some pre-made Ufunc loops:
PyUfunc_f_f |
float elementwise_func(float input_1) |
PyUfunc_ff_f |
float elementwise_func(float input_1, float input_2) |
PyUfunc_d_d |
double elementwise_func(double input_1) |
PyUfunc_dd_d |
double elementwise_func(double input_1, double input_2) |
PyUfunc_D_D |
elementwise_func(complex_double *input, complex_double* output) |
PyUfunc_DD_D |
elementwise_func(complex_double *in1, complex_double *in2, complex_double* out) |
Type codes:
NPY_BOOL, NPY_BYTE, NPY_UBYTE, NPY_SHORT, NPY_USHORT, NPY_INT, NPY_UINT,
NPY_LONG, NPY_ULONG, NPY_LONGLONG, NPY_ULONGLONG, NPY_FLOAT, NPY_DOUBLE,
NPY_LONGDOUBLE, NPY_CFLOAT, NPY_CDOUBLE, NPY_CLONGDOUBLE, NPY_DATETIME,
NPY_TIMEDELTA, NPY_OBJECT, NPY_STRING, NPY_UNICODE, NPY_VOID
2.2.2.3. Solution: building an ufunc from scratch¶
# The elementwise function
# ------------------------
cdef void mandel_single_point(double complex *z_in,
double complex *c_in,
double complex *z_out) nogil:
#
# The Mandelbrot iteration
#
#
# Some points of note:
#
# - It's *NOT* allowed to call any Python functions here.
#
# The Ufunc loop runs with the Python Global Interpreter Lock released.
# Hence, the ``nogil``.
#
# - And so all local variables must be declared with ``cdef``
#
# - Note also that this function receives *pointers* to the data;
# the "traditional" solution to passing complex variables around
#
cdef double complex z = z_in[0]
cdef double complex c = c_in[0]
cdef int k # the integer we use in the for loop
# Straightforward iteration
for k in range(100):
z = z*z + c
if z.real**2 + z.imag**2 > 1000:
break
# Return the answer for this point
z_out[0] = z
# Boilerplate Cython definitions
#
# Pulls definitions from the Numpy C headers.
# -------------------------------------------
from numpy cimport import_array, import_ufunc
from numpy cimport (PyUFunc_FromFuncAndData,
PyUFuncGenericFunction)
from numpy cimport NPY_CDOUBLE
from numpy cimport PyUFunc_DD_D
# Required module initialization
# ------------------------------
import_array()
import_ufunc()
# The actual ufunc declaration
# ----------------------------
cdef PyUFuncGenericFunction loop_func[1]
cdef char input_output_types[3]
cdef void *elementwise_funcs[1]
loop_func[0] = PyUFunc_DD_D
input_output_types[0] = NPY_CDOUBLE
input_output_types[1] = NPY_CDOUBLE
input_output_types[2] = NPY_CDOUBLE
elementwise_funcs[0] = <void*>mandel_single_point
mandel = PyUFunc_FromFuncAndData(
loop_func,
elementwise_funcs,
input_output_types,
1, # number of supported input types
2, # number of input args
1, # number of output args
0, # `identity` element, never mind this
"mandel", # function name
"mandel(z, c) -> computes iterated z*z + c", # docstring
0 # unused
)
"""
Plot Mandelbrot
================
Plot the Mandelbrot ensemble.
"""
import numpy as np
import mandel
x = np.linspace(-1.7, 0.6, 1000)
y = np.linspace(-1.4, 1.4, 1000)
c = x[None,:] + 1j*y[:,None]
z = mandel.mandel(c, c)
import matplotlib.pyplot as plt
plt.imshow(abs(z)**2 < 1000, extent=[-1.7, 0.6, -1.4, 1.4])
plt.gray()
plt.show()
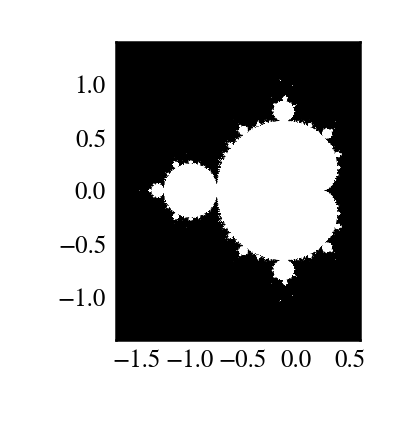
Note
Most of the boilerplate could be automated by these Cython modules:
Several accepted input types
例如,支持单精度和双精度版本
cdef void mandel_single_point(double complex *z_in,
double complex *c_in,
double complex *z_out) nogil:
...
cdef void mandel_single_point_singleprec(float complex *z_in,
float complex *c_in,
float complex *z_out) nogil:
...
cdef PyUFuncGenericFunction loop_funcs[2]
cdef char input_output_types[3*2]
cdef void *elementwise_funcs[1*2]
loop_funcs[0] = PyUFunc_DD_D
input_output_types[0] = NPY_CDOUBLE
input_output_types[1] = NPY_CDOUBLE
input_output_types[2] = NPY_CDOUBLE
elementwise_funcs[0] = <void*>mandel_single_point
loop_funcs[1] = PyUFunc_FF_F
input_output_types[3] = NPY_CFLOAT
input_output_types[4] = NPY_CFLOAT
input_output_types[5] = NPY_CFLOAT
elementwise_funcs[1] = <void*>mandel_single_point_singleprec
mandel = PyUFunc_FromFuncAndData(
loop_func,
elementwise_funcs,
input_output_types,
2, # number of supported input types <----------------
2, # number of input args
1, # number of output args
0, # `identity` element, never mind this
"mandel", # function name
"mandel(z, c) -> computes iterated z*z + c", # docstring
0 # unused
)
2.2.2.4. Generalized ufuncs¶
ufunc
output = elementwise_function(input)Both
outputandinputcan be a single array element only.
generalized ufunc
outputandinputcan be arrays with a fixed number of dimensions例如,矩阵的积(对角元素的总和):
input shape = (n, n) output shape = () i.e. scalar (n, n) -> ()矩阵的积:
input_1 shape = (m, n) input_2 shape = (n, p) output shape = (m, p) (m, n), (n, p) -> (m, p)
- This is called the “signature” of the generalized ufunc
- The dimensions on which the g-ufunc acts, are “core dimensions”
Status in NumPy
g-ufuncs are in NumPy already ...
new ones can be created with
PyUFunc_FromFuncAndDataAndSignaturemost linear-algebra functions are implemented as g-ufuncs to enable working with stacked arrays:
>>> import numpy as np >>> np.linalg.det(np.random.rand(3, 5, 5)) array([ 0.00965823, -0.13344729, 0.04583961]) >>> np.linalg._umath_linalg.det.signature '(m,m)->()'
we also ship with a few g-ufuncs for testing, ATM:
>>> import numpy.core.umath_tests as ut >>> ut.matrix_multiply.signature '(m,n),(n,p)->(m,p)' >>> x = np.ones((10, 2, 4)) >>> y = np.ones((10, 4, 5)) >>> ut.matrix_multiply(x, y).shape (10, 2, 5)
in both examples the last two dimensions became core dimensions, and are modified as per the signature
otherwise, the g-ufunc operates “elementwise”
matrix multiplication this way could be useful for operating on many small matrices at once
Generalized ufunc loop
Matrix multiplication (m,n),(n,p) -> (m,p)
void gufunc_loop(void **args, int *dimensions, int *steps, void *data)
{
char *input_1 = (char*)args[0]; /* these are as previously */
char *input_2 = (char*)args[1];
char *output = (char*)args[2];
int input_1_stride_m = steps[3]; /* strides for the core dimensions */
int input_1_stride_n = steps[4]; /* are added after the non-core */
int input_2_strides_n = steps[5]; /* steps */
int input_2_strides_p = steps[6];
int output_strides_n = steps[7];
int output_strides_p = steps[8];
int m = dimension[1]; /* core dimensions are added after */
int n = dimension[2]; /* the main dimension; order as in */
int p = dimension[3]; /* signature */
int i;
for (i = 0; i < dimensions[0]; ++i) {
matmul_for_strided_matrices(input_1, input_2, output,
strides for each array...);
input_1 += steps[0];
input_2 += steps[1];
output += steps[2];
}
}
2.2.3. Interoperability features¶
2.2.3.1. Sharing multidimensional, typed data¶
Suppose you
- Write a library than handles (multidimensional) binary data,
- Want to make it easy to manipulate the data with NumPy, or whatever other library,
- ... but would not like to have NumPy as a dependency.
Currently, 3 solutions:
- the “old” buffer interface
- the array interface
- the “new” buffer interface (PEP 3118)
2.2.3.2. The old buffer protocol¶
- Only 1-D buffers
- No data type information
- C-level interface;
PyBufferProcs tp_as_bufferin the type object - But it’s integrated into Python (e.g. strings support it)
Mini-exercise using Pillow (Python Imaging Library):
See also
pilbuffer.py
>>> from PIL import Image
>>> data = np.zeros((200, 200, 4), dtype=np.int8)
>>> data[:, :] = [255, 0, 0, 255] # Red
>>> # In PIL, RGBA images consist of 32-bit integers whose bytes are [RR,GG,BB,AA]
>>> data = data.view(np.int32).squeeze()
>>> img = Image.frombuffer("RGBA", (200, 200), data, "raw", "RGBA", 0, 1)
>>> img.save('test.png')
Q:
Check what happens ifdatais now modified, andimgsaved again.
2.2.3.3. The old buffer protocol¶
"""
From buffer
============
Show how to exchange data between numpy and a library that only knows
the buffer interface.
"""
import numpy as np
import Image
# Let's make a sample image, RGBA format
x = np.zeros((200, 200, 4), dtype=np.int8)
x[:,:,0] = 254 # red
x[:,:,3] = 255 # opaque
data = x.view(np.int32) # Check that you understand why this is OK!
img = Image.frombuffer("RGBA", (200, 200), data)
img.save('test.png')
#
# Modify the original data, and save again.
#
# It turns out that PIL, which knows next to nothing about Numpy,
# happily shares the same data.
#
x[:,:,1] = 254
img.save('test2.png')


2.2.3.4. Array interface protocol¶
- Multidimensional buffers
- Data type information present
- NumPy-specific approach; slowly deprecated (but not going away)
- Not integrated in Python otherwise
See also
Documentation: http://docs.scipy.org/doc/numpy/reference/arrays.interface.html
>>> x = np.array([[1, 2], [3, 4]])
>>> x.__array_interface__
{'data': (171694552, False), # memory address of data, is readonly?
'descr': [('', '<i4')], # data type descriptor
'typestr': '<i4', # same, in another form
'strides': None, # strides; or None if in C-order
'shape': (2, 2),
'version': 3,
}
- ::
>>> from PIL import Image >>> img = Image.open('data/test.png') >>> img.__array_interface__ {'data': ..., 'shape': (200, 200, 4), 'typestr': '|u1'} >>> x = np.asarray(img) >>> x.shape (200, 200, 4)
Note
A more C-friendly variant of the array interface is also defined.
2.2.4. Array siblings: chararray, maskedarray, matrix¶
2.2.4.1. chararray: vectorized string operations¶
>>> x = np.array(['a', ' bbb', ' ccc']).view(np.chararray)
>>> x.lstrip(' ')
chararray(['a', 'bbb', 'ccc'],
dtype='...')
>>> x.upper()
chararray(['A', ' BBB', ' CCC'],
dtype='...')
Note
.view() has a second meaning: it can make an ndarray an instance of a specialized ndarray subclass
2.2.4.2. masked_array missing data¶
Masked arrays are arrays that may have missing or invalid entries.
For example, suppose we have an array where the fourth entry is invalid:
>>> x = np.array([1, 2, 3, -99, 5])
One way to describe this is to create a masked array:
>>> mx = np.ma.masked_array(x, mask=[0, 0, 0, 1, 0])
>>> mx
masked_array(data = [1 2 3 -- 5],
mask = [False False False True False],
fill_value = 999999)
Masked mean ignores masked data:
>>> mx.mean()
2.75
>>> np.mean(mx)
2.75
Warning
Not all NumPy functions respect masks, for instance np.dot, so check the return types.
The masked_array returns a view to the original array:
>>> mx[1] = 9
>>> x
array([ 1, 9, 3, -99, 5])
2.2.4.2.1. The mask¶
You can modify the mask by assigning:
>>> mx[1] = np.ma.masked
>>> mx
masked_array(data = [1 -- 3 -- 5],
mask = [False True False True False],
fill_value = 999999)
The mask is cleared on assignment:
>>> mx[1] = 9
>>> mx
masked_array(data = [1 9 3 -- 5],
mask = [False False False True False],
fill_value = 999999)
The mask is also available directly:
>>> mx.mask
array([False, False, False, True, False], dtype=bool)
The masked entries can be filled with a given value to get an usual array back:
>>> x2 = mx.filled(-1)
>>> x2
array([ 1, 9, 3, -1, 5])
The mask can also be cleared:
>>> mx.mask = np.ma.nomask
>>> mx
masked_array(data = [1 9 3 -99 5],
mask = [False False False False False],
fill_value = 999999)
2.2.4.2.2. Domain-aware functions¶
The masked array package also contains domain-aware functions:
>>> np.ma.log(np.array([1, 2, -1, -2, 3, -5]))
masked_array(data = [0.0 0.6931471805599453 -- -- 1.0986122886681098 --],
mask = [False False True True False True],
fill_value = 1e+20)
Note
Streamlined and more seamless support for dealing with missing data in arrays is making its way into NumPy 1.7. Stay tuned!
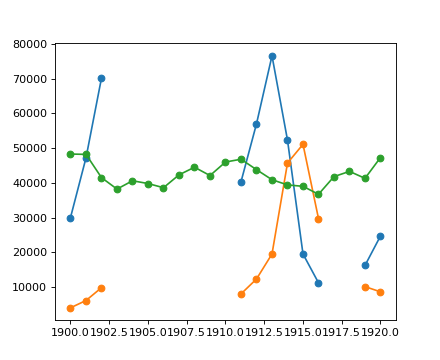
Example: Masked statistics
Canadian rangers were distracted when counting hares and lynxes in 1903-1910 and 1917-1918, and got the numbers are wrong. (Carrot farmers stayed alert, though.) Compute the mean populations over time, ignoring the invalid numbers.
>>> data = np.loadtxt('data/populations.txt')
>>> populations = np.ma.masked_array(data[:,1:])
>>> year = data[:, 0]
>>> bad_years = (((year >= 1903) & (year <= 1910))
... | ((year >= 1917) & (year <= 1918)))
>>> # '&' means 'and' and '|' means 'or'
>>> populations[bad_years, 0] = np.ma.masked
>>> populations[bad_years, 1] = np.ma.masked
>>> populations.mean(axis=0)
masked_array(data = [40472.72727272727 18627.272727272728 42400.0],
mask = [False False False],
fill_value = 1e+20)
>>> populations.std(axis=0)
masked_array(data = [21087.656489006717 15625.799814240254 3322.5062255844787],
mask = [False False False],
fill_value = 1e+20)
Note that Matplotlib knows about masked arrays:
>>> plt.plot(year, populations, 'o-')
[<matplotlib.lines.Line2D object at ...>, ...]
[source code, hires.png, pdf]
2.2.4.3. recarray: purely convenience¶
>>> arr = np.array([('a', 1), ('b', 2)], dtype=[('x', 'S1'), ('y', int)])
>>> arr2 = arr.view(np.recarray)
>>> arr2.x
chararray(['a', 'b'],
dtype='|S1')
>>> arr2.y
array([1, 2])
2.2.4.4. matrix: convenience?¶
- always 2-D
*is the matrix product, not the elementwise one
>>> np.matrix([[1, 0], [0, 1]]) * np.matrix([[1, 2], [3, 4]])
matrix([[1, 2],
[3, 4]])
2.2.5. Summary¶
- Anatomy of the ndarray: data, dtype, strides.
- Universal functions: elementwise operations, how to make new ones
- Ndarray subclasses
- Various buffer interfaces for integration with other tools
- Recent additions: PEP 3118, generalized ufuncs
2.2.6. 贡献NumPy/Scipy ¶
Get this tutorial: http://www.euroscipy.org/talk/882
2.2.6.1. Why¶
- “There’s a bug?”
- “I don’t understand what this is supposed to do?”
- “I have this fancy code. Would you like to have it?”
- “I’d like to help! What can I do?”
2.2.6.2. Reporting bugs¶
- Bug tracker (prefer this)
- http://projects.scipy.org/numpy
- http://projects.scipy.org/scipy
- Click the “Register” link to get an account
- Mailing lists ( scipy.org/Mailing_Lists )
- If you’re unsure
- No replies in a week or so? Just file a bug ticket.
2.2.6.2.1. Good bug report¶
Title: numpy.random.permutations fails for non-integer arguments
I'm trying to generate random permutations, using numpy.random.permutations
When calling numpy.random.permutation with non-integer arguments
it fails with a cryptic error message::
>>> np.random.permutation(12)
array([11, 5, 8, 4, 6, 1, 9, 3, 7, 2, 10, 0])
>>> np.random.permutation(12.)
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "mtrand.pyx", line 3311, in mtrand.RandomState.permutation
File "mtrand.pyx", line 3254, in mtrand.RandomState.shuffle
TypeError: len() of unsized object
This also happens with long arguments, and so
np.random.permutation(X.shape[0]) where X is an array fails on 64
bit windows (where shape is a tuple of longs).
It would be great if it could cast to integer or at least raise a
proper error for non-integer types.
I'm using NumPy 1.4.1, built from the official tarball, on Windows
64 with Visual studio 2008, on Python.org 64-bit Python.
What are you trying to do?
Small code snippet reproducing the bug (if possible)
- What actually happens
- What you’d expect
Platform (Windows / Linux / OSX, 32/64 bits, x86/PPC, ...)
Version of NumPy/Scipy
>>> print(np.__version__) 1...
Check that the following is what you expect
>>> print(np.__file__) /...
In case you have old/broken NumPy installations lying around.
If unsure, try to remove existing NumPy installations, and reinstall...
2.2.6.3. Contributing to documentation¶
Documentation editor
Registration
Register an account
Subscribe to
scipy-devmailing list (subscribers-only)Problem with mailing lists: you get mail
But: you can turn mail delivery off
“change your subscription options”, at the bottom of
Send a mail @
scipy-devmailing list; ask for activation:To: scipy-dev@scipy.org Hi, I'd like to edit NumPy/Scipy docstrings. My account is XXXXX Cheers, N. N.
- Check the style guide:
- http://docs.scipy.org/doc/numpy/
- Don’t be intimidated; to fix a small thing, just fix it
- Edit
Edit sources and send patches (as for bugs)
Complain on the mailing list
2.2.6.4. Contributing features¶
Ask on mailing list, if unsure where it should go
Write a patch, add an enhancement ticket on the bug tracket
OR, create a Git branch implementing the feature + add enhancement ticket.
- Especially for big/invasive additions
- http://projects.scipy.org/numpy/wiki/GitMirror
- http://www.spheredev.org/wiki/Git_for_the_lazy
# Clone numpy repository git clone --origin svn http://projects.scipy.org/git/numpy.git numpy cd numpy # Create a feature branch git checkout -b name-of-my-feature-branch svn/trunk <edit stuff> git commit -a
- Create account on https://github.com (or anywhere)
- Create a new repository @ Github
- Push your work to github
git remote add github git@github:USERNAME/REPOSITORYNAME.git git push github name-of-my-feature-branch
2.2.6.5. How to help, in general¶
Bug fixes always welcome!
- What irks you most
- Browse the tracker
Documentation work
API docs: improvements to docstrings
- Know some Scipy module well?
User guide
Needs to be done eventually.
Want to think? Come up with a Table of Contents
Ask on communication channels:
numpy-discussionlistscipy-devlist
